RBFE¶
RBFE API is in development
RBFE functionality is still under development.
Removed: Complex.rbfe / deeporigin.drug_discovery.rbfe
The RBFE module and Complex.rbfe are no longer in the SDK. This tutorial is historical (legacy Complex orchestration). Prefer SystemPrep and platform RBFE tools for new work.
This document describes how to run a RBFE simulation using Deep Origin tools.
Prerequisites¶
We assume that we have an initialized and configured Complex object:
from deeporigin.drug_discovery import Complex, BRD_DATA_DIR
sim = Complex.from_dir(BRD_DATA_DIR)
Starting an RBFE run¶
Single pair of ligands¶
To run an end-to-end RBFE workflow on a single pair of ligands, we use:
job = sim.rbfe.run_ligand_pair(
ligand1=sim.ligands[0], # for example
ligand2=sim.ligands[1],
)
This queues up a task on Deep Origin. When it completes, outputs will be written to the appropriate column in this database.
You will see a message printed to screen similar to:
Expected output
🧬 Job started with ID: 20f05e96, execution ID: x9rl5eghrpqwyiciehc3e
Constructing a network¶
Constructing a network
View the documentation for LigandSet to learn how to construct a network.
Run an entire network¶
To run an entire network, use:
job = sim.rbfe.run_network()
This creates a compound job on Deep Origin, where each ligand pair will be run in parallel.
Parameters¶
The end to end RBFE tool has a number of user-accessible parameters. To view all parameters, use:
from deeporigin.drug_discovery import Complex, BRD_DATA_DIR
sim = Complex.from_dir(BRD_DATA_DIR)
sim.rbfe._params["end_to_end"]
Expected output
This will print a dictionary of the parameters used for RBFE, similar to:
{
"complex_prep": {
"include_ligands": 1,
"include_protein": 1,
"sysprep_params": {
"charge_method": "bcc",
"do_loop_modelling": false,
"force_field": "ff14SB",
"is_lig_protonated": false,
"is_protein_protonated": false,
"keep_waters": false,
"lig_force_field": "gaff2",
"padding": 1.0,
"save_gmx_files": false
},
"test_run": 0,
"thread_pinning": 1,
"thread_pinning_offset": 0
},
"ligand1_prep": {
"include_ligands": 1,
"include_protein": 0,
"sysprep_params": {
"charge_method": "bcc",
"do_loop_modelling": false,
"force_field": "ff14SB",
"is_lig_protonated": false,
"is_protein_protonated": false,
"keep_waters": false,
"lig_force_field": "gaff2",
"padding": 1.0,
"save_gmx_files": false
},
"test_run": 0,
"thread_pinning": 1,
"thread_pinning_offset": 0
},
"ligand2_prep": {
"include_ligands": 1,
"include_protein": 0,
"sysprep_params": {
"charge_method": "bcc",
"do_loop_modelling": false,
"force_field": "ff14SB",
"is_lig_protonated": false,
"is_protein_protonated": false,
"keep_waters": false,
"lig_force_field": "gaff2",
"padding": 1.0,
"save_gmx_files": false
},
"test_run": 0,
"thread_pinning": 1,
"thread_pinning_offset": 0
},
"rbfe": {
"add_fep_repeats": 0,
"amend": "__NO_AMEND",
"annihilate": true,
"atom_mapping_threshold": 0.01,
"em_all": true,
"em_solvent": true,
"emeq_md_options": {
"T": 298.15,
"cutoff": 0.9,
"fourier_spacing": 0.12,
"hydrogen_mass": 2.0,
"Δt": 0.004
},
"fep_windows": [
{
"restraints_A": [
0.0,
0.01,
0.025,
0.05,
0.1,
0.35,
0.5,
0.75,
1.0
]
},
{
"coul_A": [
1.0,
0.8,
0.6,
0.4,
0.2,
0.0
]
},
{
"vdw_A": [
1.0,
0.9,
0.8,
0.7,
0.6,
0.5,
0.4,
0.3,
0.2,
0.1,
0.0
]
}
],
"mbar": 1,
"npt_reduce_restraints_ns": 2.0,
"nvt_heating_ns": 1.0,
"prod_md_options": {
"T": 298.15,
"barostat": "MonteCarloBarostat",
"barostat_exchange_interval": 500,
"cutoff": 0.9,
"fourier_spacing": 0.12,
"hydrogen_mass": 2.0,
"integrator": "BAOABIntegrator",
"Δt": 0.004
},
"repeats": 1,
"run_name": "binding",
"skip_emeq": "__NO",
"softcore_alpha": 0.5,
"steps": 1250000,
"system": "complex",
"test_run": 0,
"thread_pinning": 1,
"thread_pinning_offset": 0,
"threads": 0,
"workers": 0
},
"solvation1": {
"add_fep_repeats": 0,
"amend": "__NO_AMEND",
"annihilate": true,
"atom_mapping_threshold": 0.01,
"em_all": true,
"em_solvent": true,
"emeq_md_options": {
"T": 298.15,
"cutoff": 0.9,
"fourier_spacing": 0.12,
"hydrogen_mass": 2.0,
"Δt": 0.004
},
"fep_windows": [
{
"coul_A": [
1.0,
0.8,
0.6,
0.4,
0.2,
0.0
]
},
{
"vdw_A": [
1.0,
0.9,
0.8,
0.7,
0.6,
0.5,
0.4,
0.3,
0.2,
0.1,
0.0
]
}
],
"mbar": 1,
"npt_reduce_restraints_ns": 0.2,
"nvt_heating_ns": 0.1,
"prod_md_options": {
"T": 298.15,
"barostat": "MonteCarloBarostat",
"barostat_exchange_interval": 500,
"cutoff": 0.9,
"fourier_spacing": 0.12,
"hydrogen_mass": 2.0,
"integrator": "BAOABIntegrator",
"Δt": 0.004
},
"repeats": 1,
"skip_emeq": "__NO",
"softcore_alpha": 0.5,
"steps": 300000,
"test_run": 0,
"thread_pinning": 1,
"thread_pinning_offset": 0,
"threads": 0,
"workers": 0
},
"solvation2": {
"add_fep_repeats": 0,
"amend": "__NO_AMEND",
"annihilate": true,
"atom_mapping_threshold": 0.01,
"em_all": true,
"em_solvent": true,
"emeq_md_options": {
"T": 298.15,
"cutoff": 0.9,
"fourier_spacing": 0.12,
"hydrogen_mass": 2.0,
"Δt": 0.004
},
"fep_windows": [
{
"coul_A": [
1.0,
0.8,
0.6,
0.4,
0.2,
0.0
]
},
{
"vdw_A": [
1.0,
0.9,
0.8,
0.7,
0.6,
0.5,
0.4,
0.3,
0.2,
0.1,
0.0
]
}
],
"mbar": 1,
"npt_reduce_restraints_ns": 0.2,
"nvt_heating_ns": 0.1,
"prod_md_options": {
"T": 298.15,
"barostat": "MonteCarloBarostat",
"barostat_exchange_interval": 500,
"cutoff": 0.9,
"fourier_spacing": 0.12,
"hydrogen_mass": 2.0,
"integrator": "BAOABIntegrator",
"Δt": 0.004
},
"repeats": 1,
"skip_emeq": "__NO",
"softcore_alpha": 0.5,
"steps": 300000,
"test_run": 0,
"thread_pinning": 1,
"thread_pinning_offset": 0,
"threads": 0,
"workers": 0
}
}
Any of these parameters are modifiable using dot notation. For example, to change the number of steps in the RBFE step, we can use:
from deeporigin.drug_discovery import Complex, BRD_DATA_DIR
sim = Complex.from_dir(BRD_DATA_DIR)
sim.rbfe._params["end_to_end"]["binding"]["steps"] = 500000
Results¶
Viewing results¶
After initiating a run, we can view results using:
sim.rbfe.show_results()
This shows a table similar to:
Expected output
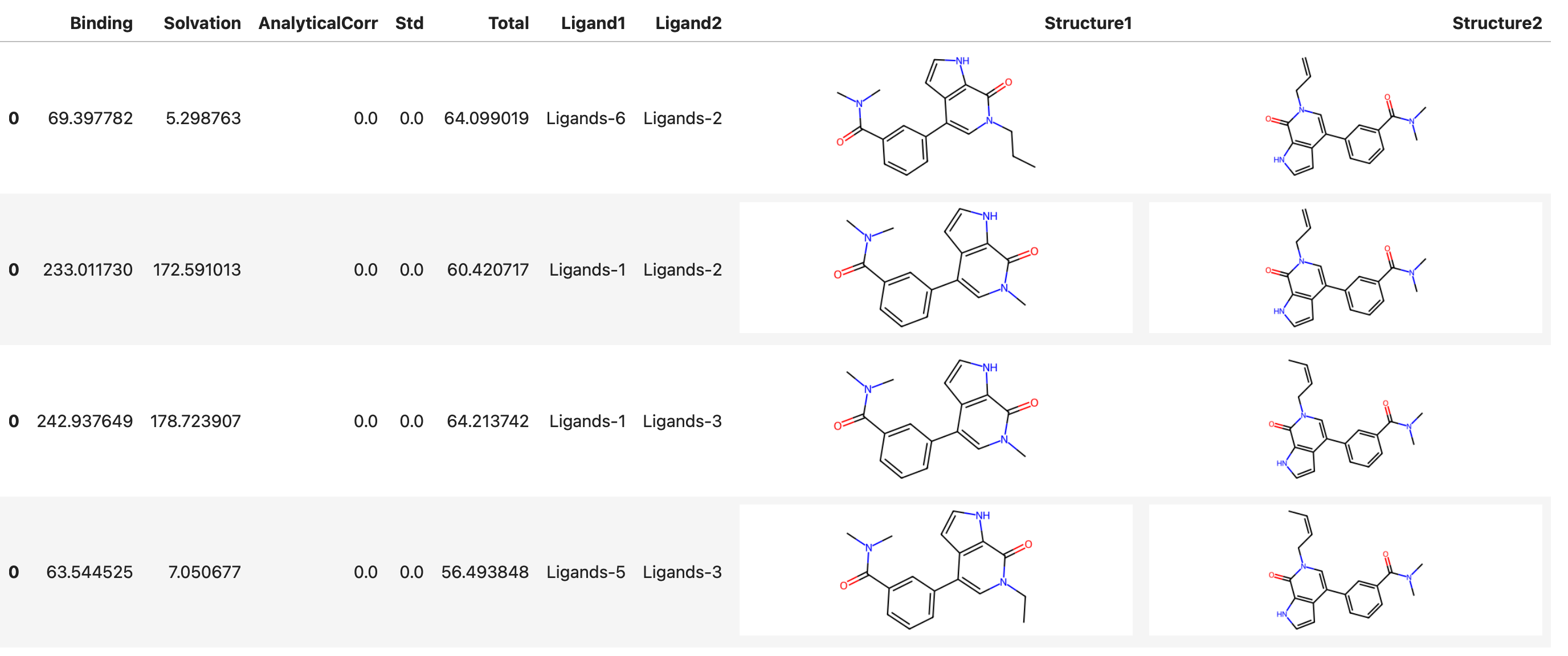
Exporting results for analysis¶
These results can be exported for analysis using:
df = sim.rbfe.get_results()
df
Expected output
| Index | Binding | Solvation | AnalyticalCorr | Std | Total | Ligand1 | Ligand2 | SMILES1 | SMILES2 |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 69.397782 | 5.298763 | 0.0 | 0.0 | 64.099019 | Ligands-6 | Ligands-2 | [H]C1=C([H])C(C(=O)N(C([H])([H])[H])C([H])([H]... | [H]C([H])=C([H])C([H])([H])N1C(=O)C2=C(C([H])=... |
| 0 | 233.01173 | 172.591013 | 0.0 | 0.0 | 60.420717 | Ligands-1 | Ligands-2 | [H]C1=C([H])C(C(=O)N(C([H])([H])[H])C([H])([H]... | [H]C([H])=C([H])C([H])([H])N1C(=O)C2=C(C([H])=... |
| 0 | 242.937649 | 178.723907 | 0.0 | 0.0 | 64.213742 | Ligands-1 | Ligands-3 | [H]C1=C([H])C(C(=O)N(C([H])([H])[H])C([H])([H]... | [H]C1=C([H])C(C(=O)N(C([H])([H])[H])C([H])([H]... |
| 0 | 63.544525 | 7.050677 | 0.0 | 0.0 | 56.493848 | Ligands-5 | Ligands-3 | [H]C1=C([H])C(C(=O)N(C([H])([H])[H])C([H])([H]... | [H]C1=C([H])C(C(=O)N(C([H])([H])[H])C([H])([H]... |